
Newsroom
A research team led by Profs. GAO Caixia and LI Jiayang from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences have engineered five saturated targeted endogenous mutagenesis editors (STEMEs) and generated de novo mutations to facilitate the directed evolution of plant genes. Their study was published in Nature Biotechnology on Jan. 13.
Heredity and variation are the basis of organismic evolution. Random mutagenesis by physical or chemical methods has long been applied to improve traits in plants, but it is labor-intensive and time-consuming.
In higher organisms, especially in plants, a target gene is usually transferred into a bacterial or yeast cell to generate the required diversity for selection, but once a target gene is no longer in situ, the functional consequences of such a change may not be the same as in the native context. Moreover, most important agronomic traits cannot be selected in bacteria or yeast.
"To establish powerful tools for directly inducing saturated targeted mutations and selection in plants will accelerate the development of agronomic traits and important functional genes," said Prof. GAO Caixia.
The researchers fused cytidine deaminase with adenosine deaminase to obtain four STEMEs. All four STEMEs efficiently produced simultaneous C>T and A>G conversions using only a sgRNA.
They also produced the fifth dual cytosine and adenine base editor - STEME-NG - to expand the targeting scope. With only 20 sgRNAs in rice protoplasts, STEME-NG can produce near-saturated mutagenesis for a 56-amino-acid portion of the rice acetyl-coenzyme A carboxylase gene (OsACC).
In a proof-of-concept experiment, the researchers used STEMEs to direct the evolution of OsACC gene in rice plants. They sprayed the regenerated rice seedlings with haloxyfop as the selection pressure. The scientists then identified three novel (P1927F, W2125C, and S1866F) and one known (W2125C) amino acid substitutions for herbicide resistance. These mutations were found to affect the haloxyfop-binding pocket directly or indirectly, based on the homology model of the CT domain of yeast ACC.
The development of STEME paves the way for directed evolution of endogenous plant genes in situ, which is important for breeding via molecular design.
Moreover, this STEME process might also be applicable beyond plants. For example, it may be useful for screening drug resistance mutations, altering cis elements on noncoding regions and correcting pathogenic SNVs in cell lines, yeast or animals.
The study was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences, the National Natural Science Foundation of China, and the National Key Research and Development Program of China.
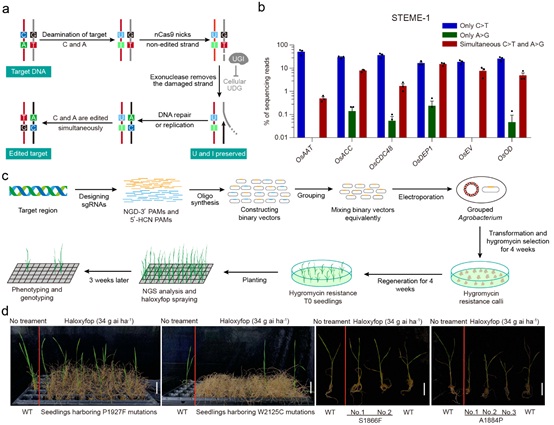
(a) STEME-mediated C >T and A >G base-editing strategy. (b) Distribution of edited DNA sequencing reads in rice protoplasts for STEME-1. (c) Procedure for mutating the OsACC CT domain via STEME using groups of individual sgRNAs. (d) Effects of STEME-induced mutations on herbicide resistance. (Image by IGDB)
