
Newsroom
Scientists from the Qingdao Institute of Bioenergy and Bioprocess Technology (QIBEBT) of the Chinese Academy of Sciences (CAS) and the Tianjin Institute of Industrial Biotechnology (TIB) of CAS, have developed a rapid, high-throughput error-correction platform that significantly improves the accuracy of synthetic DNA. The system, known as engineered MutS-based magnetic bead separation (eMBS), offers an automated approach to removing error-containing DNA.
The study was published in Synthetic and Systems Biotechnology on March 6.
High-fidelity DNA synthesis is essential for synthetic biology, especially for constructing long DNA sequences, large libraries, and automated workflows. However, widely used methods such as high-performance liquid chromatography (HPLC) and polyacrylamide gel electrophoresis (PAGE) are often labor-intensive and difficult to scale. The eMBS platform addresses these challenges by offering a faster, more selective strategy for error correction.
The system is built around an engineered Escherichia coli MutS protein, with a key mutation (E38N) that improves mismatch recognition while minimizing non-specific binding. Two cysteine substitutions (L157C and G233C) were introduced to enhance protein stability and affinity for errors involving insertions, deletions, and substitutions. The fusion protein is immobilized on streptavidin-coated magnetic beads, enabling selective capture of error-containing heteroduplexes, while leaving the correct DNA in the solution for recovery.
The full eMBS workflow takes just 20 minutes, including 10 minutes for binding, five minutes for magnetic bead capture, and five minutes for separation and recovery. Unlike traditional methods, eMBS avoids electrophoresis and column purification, thereby reducing hands-on time and sample loss. The platform can process up to 96 samples simultaneously, offering a substantial throughput advantage over traditional approaches.
Validation experiments demonstrated that eMBS reduced total DNA synthesis errors by 13.1% to 63.3% across substrate pools with initial correct sequence rates ranging from 73.6% to 98.6%. Under optimal conditions, the platform achieved an error-free DNA ratio of 99.1%. The engineered E38N-containing variant performed particularly well, especially in correcting insertion and deletion errors.
"This platform represents a breakthrough in improving the fidelity and scalability of DNA synthesis," said Prof. ZHANG Jia from the Single-Cell Center of QIBEBT, corresponding author of the the study. Co-author Prof. SHENG Xiang from TIB added that eMBS offers a practical solution for high-fidelity DNA synthesis in automated workflows, making it a valuable tool for both research and industrial-scale applications.
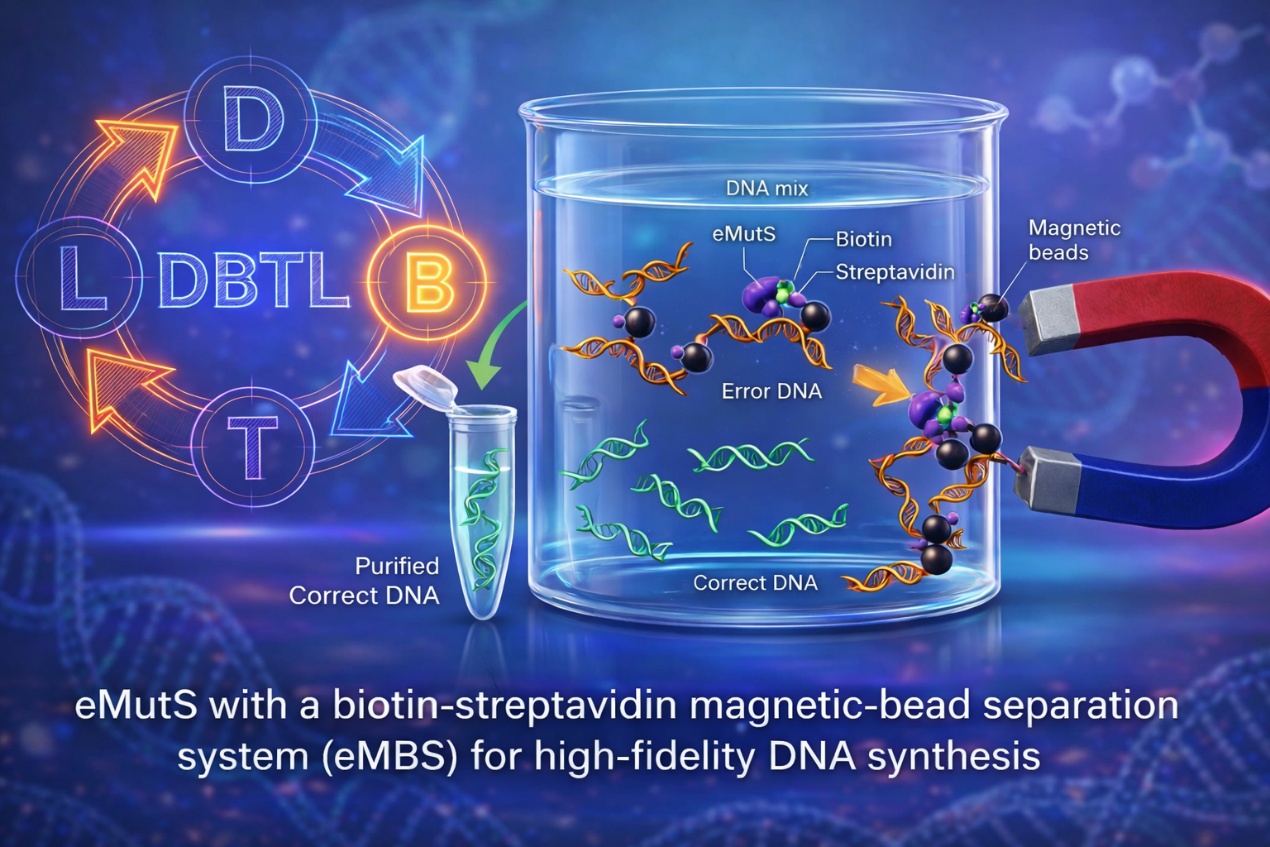
eMutS with a biotin-streptavidin magnetic-bead separationsystem (eMBS) for high-fidelity DNA synthesis (Image by LIU Yang and SHU Yang)
WANG Xiaohang
Qingdao Institute of Bioenergy and Bioprocess Technology
E-mail: wangxiaohang@qibebt.ac.cn
